Coarse-graining Tutorial¶
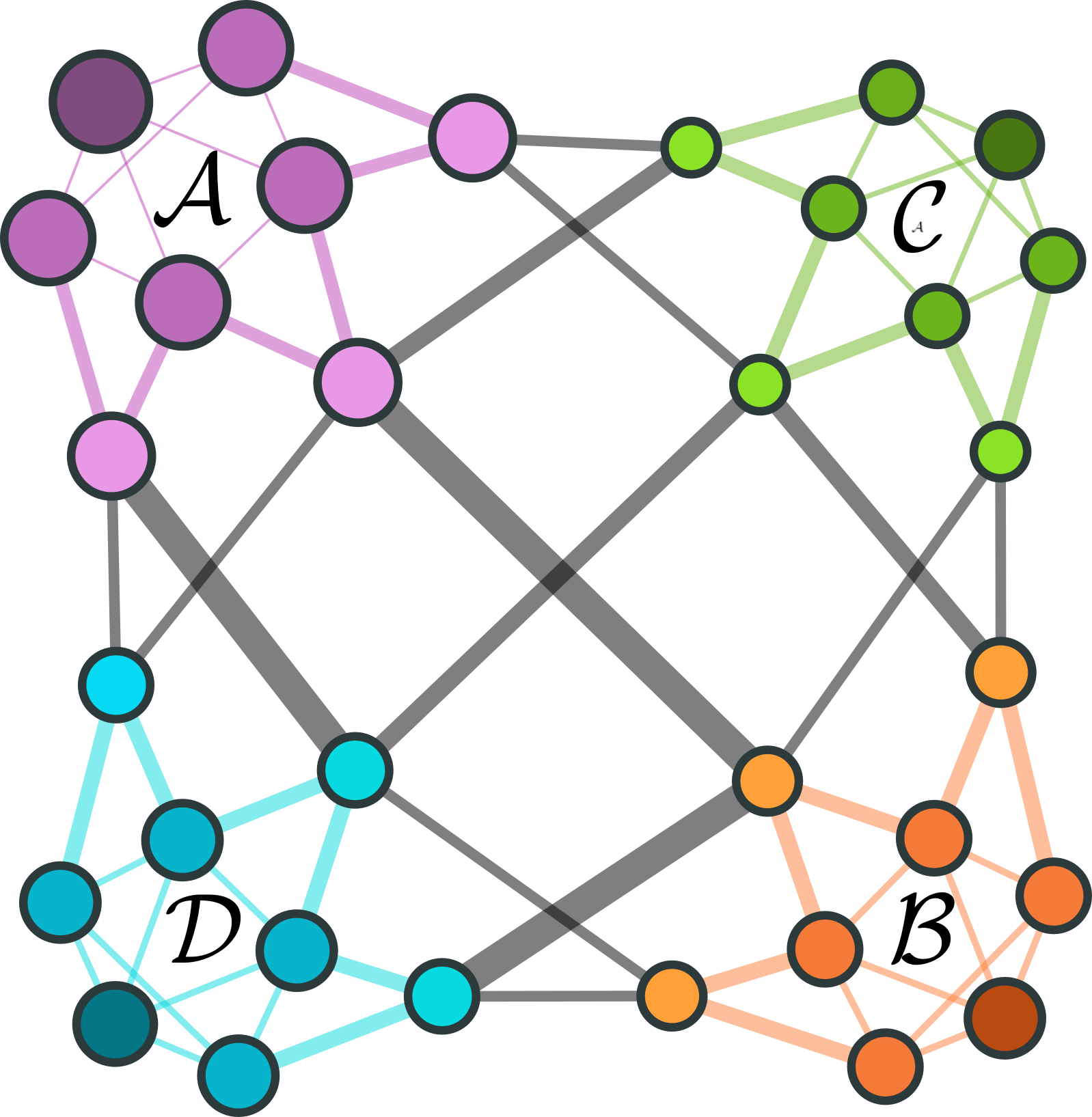
We illustrate the tools avalable in the PyGT package for the dimensionality reduction of Markov chains using a model 32-state network. The network can be divided into 4 competing macrostates. We will compute the optimal \(4 \times 4\) coarse-grained rate matrix with various numerical methods and compare the reduced dynamics to the original model.
[2]:
#uncomment if PyGT not installed via pip
import sys; sys.path.insert(0,"../")
import PyGT
#other modules
import numpy as np
import scipy.linalg as spla
from scipy.sparse import issparse, diags
from pathlib import Path
import pandas as pd
from matplotlib import pyplot as plt
# optional
try:
import seaborn as sns
sns.set()
has_seaborn=True
except:
has_seaborn=False
Model 32-state network¶
Each community has 8 nodes, including 1 attractor node, 4 internal nodes, and 3 boundary nodes. Nodes are colored by the community to which they belong. Darker, larger nodes have higher equilbrium occupation probabilities, and thicker edges indicate slower transitions.
[3]:
from IPython.display import Image
Image(filename = "32state.png", width = 300)
GT setup¶
Let’s load in the Markov chain as well as its community structure. Community assignments are specified in a single-column file where each line contains the community ID of the node corresponding to the line number.
[4]:
data_path = Path('KTN_data/32state')
temp = 10.0
beta = 1./temp
#GT setup
B, K, tau, N, u, s, Emin, retained = PyGT.io.load_ktn(path=data_path,beta=beta)
#rate matrix with columns that sum to zero
# K has no diagonal entries
if issparse(K):
Q = K - diags(1.0/tau)
else:
Q = K - np.diag(1.0/tau)
BF = beta*u-s
BF -= BF.min()
#stationary distribution
pi = np.exp(-BF)
pi /= pi.sum()
#A and B sets
AS,BS = PyGT.io.load_ktn_AB(data_path,retained)
#Read in community structure
comms = PyGT.io.read_communities(data_path/'communities.dat', retained, screen=True)
for comm in comms:
if np.all(comms[comm] == AS):
print(f'Community A: {comm}')
if np.all(comms[comm] == BS):
print(f'Community B: {comm}')
Community 0: 8
Community 1: 8
Community 2: 8
Community 3: 8
Community A: 0
Community B: 3
Matrix of inter-microstate MFPTs with GT vs. linear algebra methods¶
The \(32 \times 32\) matrix of inter-microstate MFPTs between all pairs of nodes can be used to obtain the optimal reduced coarse-grained Markov chain for a given community structure. Let’s compute this matrix with GT and with two alternative linear algebra methods: inversion to obtain the fundamental matrix and solving a linear equation.
[5]:
#compute matrix of inter-microstate MFPTs with GT
mfpt_gt = PyGT.mfpt.full_MFPT_matrix(B, tau)
#check that the Kemeny constant is indeed constant
kemeny, success = PyGT.tools.check_kemeny(pi, mfpt_gt)
if success:
print("Kemeny constant from mfpts with GT: ", kemeny)
#compute matrix of inter-microstate MFPTs with fundamental matrix
ktn = PyGT.tools.Analyze_KTN(data_path, K=Q.todense(), pi=pi, commdata='communities.dat')
mfpt_fund = ktn.get_intermicrostate_mfpts_fundamental_matrix()
kemeny_fund, success = PyGT.tools.check_kemeny(pi, mfpt_fund)
if success:
print("Kemeny constant from mfpts with fundamental matrix: ", kemeny_fund)
#compute matrix of inter-microstate MFPTs by solving a linear equation
mfpt_lin = ktn.get_intermicrostate_mfpts_linear_solve()
kemeny_lin, success = PyGT.tools.check_kemeny(pi, mfpt_lin)
if success:
print("Kemeny constant from mfpts with linear solve: ", kemeny_lin)
Kemeny constant from mfpts with GT: 97.53002029348983
Kemeny constant from mfpts with fundamental matrix: 97.53002029348986
Kemeny constant from mfpts with linear solve: 97.53002029348977
Compute inter-community weighted-MFPTs¶
[6]:
#compute weighted-MFPT between communities
commpi = ktn.get_comm_stat_probs(np.log(pi), log=False)
ktn.commpi = commpi
ncomms = len(commpi)
pt = ktn.get_intercommunity_weighted_MFPTs(mfpt_gt)
#Kemeny constant of reduced Markov chain
print("Weighted-MFPT matrix:")
print(pt)
c_kemeny, success = PyGT.tools.check_kemeny(commpi, pt)
if success:
print('\nKemeny constant of coarse-grained Markov chain: ', c_kemeny)
Weighted-MFPT matrix:
[[ 0. 53.8206528 54.95852132 69.11554233]
[53.00851035 0. 66.57240778 52.25954611]
[54.71235842 67.13838734 0. 54.16914917]
[69.93796797 53.8941142 55.2377377 0. ]]
Kemeny constant of coarse-grained Markov chain: 44.06002077112634
Different routes to obtain the optimal coarse-grained CTMC¶
In Kannan et al. J. Chem. Phys. (2020), we discuss three different expression for the optimal coarse-grained rate matrix given a partitioning of the \(V\) nodes in the original Markov chain into \(N\) communities: the HS relation, the KKRA relation, and an expression obtained from inverting the matrix of weighted-MFPTs. We illustrate the computation of all 3 methods below:
[7]:
""" Three different version of the optimal reduced CTMC."""
#1) the original HS relation
K_hs = ktn.construct_coarse_rate_matrix_Hummer_Szabo()
#2) the KKRA relation involving inversion of matrix of inter-microstate mfpts
K_kkra = ktn.construct_coarse_rate_matrix_KKRA(GT=True)
#3) based on inversion of weighted-MFPTs
K_invert = spla.inv(pt)@(np.diag(1./commpi) - np.ones((ncomms,ncomms)))
print('Optimal reduced CTMC from Hummer-Szabo relation:')
print(K_hs)
print('Optimal reduced CTMC from KKRA relation:')
print(K_kkra)
print('Optimal reduced CTMC from inversion of weighted-MFPT matrix:')
print(K_invert)
#check that detailed balance is satisfied
if not PyGT.tools.check_detailed_balance(commpi, K_invert):
print('Detailed balance not satisfied for K_C.')
if not PyGT.tools.check_detailed_balance(pi, Q):
print('Detailed balance not satisfied for K')
Optimal reduced CTMC from Hummer-Szabo relation:
[[-0.05239043 0.02620963 0.02440272 0.0036629 ]
[ 0.02493137 -0.05761829 0.00478585 0.02595583]
[ 0.02390442 0.00492849 -0.05366994 0.0247118 ]
[ 0.00355464 0.02648017 0.02448137 -0.05433052]]
Optimal reduced CTMC from KKRA relation:
[[-0.05239043 0.02620963 0.02440272 0.0036629 ]
[ 0.02493137 -0.05761829 0.00478585 0.02595583]
[ 0.02390442 0.00492849 -0.05366994 0.0247118 ]
[ 0.00355464 0.02648017 0.02448137 -0.05433052]]
Optimal reduced CTMC from inversion of weighted-MFPT matrix:
[[-0.05239043 0.02620963 0.02440272 0.0036629 ]
[ 0.02493137 -0.05761829 0.00478585 0.02595583]
[ 0.02390442 0.00492849 -0.05366994 0.0247118 ]
[ 0.00355464 0.02648017 0.02448137 -0.05433052]]
Numerical comparison of coarse-grained Markov chains¶
To compare the numerical stability of these various routes to obtain the optimal reduced CTMC, let’s compute the mean first passage times \(\mathcal{A} \leftrightarrow \mathcal{B}\) on the original network and compare it to the corresponding observables on the various reduced networks.
[8]:
def compare_HS_LEA(temps, data_path):
""" Calculate coarse-grained rate matrices using the 3 versions of the optimal
reudced Markov chain and the local equilibrium approximation (LEA).
Compute MFPTAB/BA on the full and coarse-grained networks. """
dfs = []
for temp in temps:
df = pd.DataFrame()
df['T'] = [temp]
#KTN input
beta = 1./temp
B, K, tau, N, u, s, Emin, retained = PyGT.io.load_ktn(path=data_path,beta=beta)
Q = K - diags(1.0/tau)
BF = beta*u-s
BF -= BF.min()
#stationary distribution
pi = np.exp(-BF)
pi /= pi.sum()
#A and B sets
AS,BS = PyGT.io.load_ktn_AB(data_path,retained)
#ktn setup
ktn = PyGT.tools.Analyze_KTN(data_path, K=Q, pi=pi, commdata='communities.dat')
commpi = ktn.commpi
ncomms = len(ktn.commpi)
#MFPT calculations on full network
full_df = PyGT.stats.compute_rates(AS, BS, B, tau, pi, fullGT=True, block=1)
df['MFPTAB'] = full_df['MFPTAB']
df['MFPTBA'] = full_df['MFPTBA']
#compute coarse-grained networks
mfpt = PyGT.mfpt.full_MFPT_matrix(B, tau)
pt = ktn.get_intercommunity_weighted_MFPTs(mfpt)
labels = []
matrices = []
try:
Rhs = ktn.construct_coarse_rate_matrix_Hummer_Szabo()
matrices.append(Rhs)
labels.append('HS')
except Exception as e:
print(f'HS had the following error: {e}')
try:
Rhs_kkra = ktn.construct_coarse_rate_matrix_KKRA(mfpt=mfpt)
matrices.append(Rhs_kkra)
labels.append('KKRA')
except Exception as e:
print(f'KKRA had the following error: {e}')
try:
Rhs_invert = spla.inv(pt)@(np.diag(1./commpi) - np.ones((ncomms,ncomms)))
matrices.append(Rhs_invert)
labels.append('PTinvert_GT')
except Exception as e:
print(f'Inversion of weighted-MFPTs from GT had the following error: {e}')
try:
Rlea = ktn.construct_coarse_rate_matrix_LEA()
matrices.append(Rlea)
labels.append('LEA')
except Exception as e:
print(f'LEA had the following error: {e}')
if len(matrices)==0:
continue
for i, R in enumerate(matrices):
""" get A->B and B->A mfpt on coarse network"""
rK = R - np.diag(np.diag(R))
escape_rates = -1*np.diag(R)
B = rK@np.diag(1./escape_rates)
tau = 1./escape_rates
#B, tau = PyGT.tools.load_CTMC(R)
Acomm = 0
Bcomm = 3
MFPTAB, MFPTBA = PyGT.mfpt.compute_MFPT(Acomm, Bcomm, B, tau, block=1)
df[f'AB_{labels[i]}'] = [MFPTAB]
df[f'BA_{labels[i]}'] = [MFPTBA]
dfs.append(df)
bigdf = pd.concat(dfs, ignore_index=True, sort=False)
return bigdf
Plot KKRA, H-S against exact, LEA and GT systems at high temperature¶
[9]:
#some mid temperature calculations
invtemps = np.linspace(0.1, 4, 6)
midtemp_df = compare_HS_LEA(1./invtemps, data_path)
[10]:
def plot_mfpts_32state(df):
"""Plot MFPTs computed on coarse-grained networks against true MFPT from full network."""
if has_seaborn:
colors = sns.color_palette("Dark2", 4)
else:
colors = ['C0','C1','C2','C3']
df.replace([np.inf, -np.inf], np.nan)
df2= df.sort_values('T')
symbols = ['-s', '--o', '-o', '--^']
rates = ['LEA', 'PTinvert_GT', 'KKRA', 'HS']
labels = rates
denom = 'MFPT'
#first plot A<-B direction
fig, (ax, ax2) = plt.subplots(1, 2, figsize=[10, 4])
ax.plot(1./df2['T'], df2['MFPTBA'], '-', color='k', label='Exact', lw=1, markersize=4)
for j, CG in enumerate(rates):
#then only plot HSK for temperatures that are not NaN
df2CG = df2[-df2[f'BA_{CG}'].isna()]
ax.plot(1./df2CG['T'], df2CG[f'BA_{CG}'],
symbols[j], label=labels[j], color=colors[j], linewidth=1,
markersize=4)
ax.set_xlabel(r'$1/T$')
ax.set_yscale('log')
ax.set_ylabel('MFPTBA')
ax.legend(frameon=True)
ax2.plot(1./df2['T'], df2['MFPTAB'], '-', color='k', label='Exact', lw=1, markersize=4)
for j, CG in enumerate(rates):
#then only plot HSK for temperatures that are not NaN
df2CG = df2[-df2[f'AB_{CG}'].isna()]
ax2.plot(1./df2CG['T'], df2CG[f'AB_{CG}'],
symbols[j], label=labels[j], color=colors[j], linewidth=1,
markersize=4)
ax2.set_xlabel(r'$1/T$')
ax2.set_ylabel('MFPTAB')
ax2.set_yscale('log')
ax2.legend(frameon=True)
fig.tight_layout()
[11]:
plot_mfpts_32state(midtemp_df)

Plot KKRA, H-S against exact, LEA and GT systems at slightly lower temperature¶
KKRA, H-S fails
[14]:
invtemps = np.linspace(5, 15, 10)
lowtemp_df = compare_HS_LEA(1./invtemps, data_path)
[16]:
plot_mfpts_32state(lowtemp_df)